Pirazinamid diren?li ve duyarl? Mycobacterium tuberculosis izolatlar?nda
pirazinamidaz yap?sal de?i?ikliklerinin ?al??mas?
Azam AHMADY1, Toktam POOLAD2, Poorya RAFEE1, Mojtaba TOUSHEH3, Manijeh KAHBAZI1,
Mohammad ARJOMANDZADEGAN1
1 Arak ?niversitesi T?p Fak?ltesi, T?berk?loz ve Pediatrik ?nfeksiyon Hastal?klar? Ara?t?rma Merkezi, Arak, ?ran,
2 ?slami Azad ?niversitesi, K?rdistan Bilim ve Ara?t?rma ?ubesi, Mikrobiyoloji B?l?m?, Tahran, ?ran,
3 ?sfahan ?niversitesi H?cresel ve Molek?ler Biyoloji B?l?m?, ?sfahan, ?ran.
?ZET
Pirazinamid diren?li ve duyarl? Mycobacterium tuberculosis izolatlar?nda pirazinamidaz yap?sal de?i?ikliklerinin ?al??mas?
Giri?: Pirazinamid, t?berk?loz tedavisindeki ilk basamak ila?lardan biridir. 359 ve 374 pnc genlerinin iki n?kleotidindeki mutasyonun, pirazinamid direnciyle y?ksek derecede ili?kili oldu?u g?sterilmi?tir.
Materyal ve Metod: Bu ?al??mada, 30 klinik Mycobacterium tuberculosis izolat?nda sekans analiziyle bu iki kodondaki mutasyonlar ara?t?r?ld?. Pirazinamid diren?li ve duyarl? izolatlarda, mutasyonlu ve mutasyonsuz bu gen taraf?ndan kodlanan protein yap?lar? ara?t?r?ld?.
Bulgular: 359 ve 374 pozisyonlar?ndaki mutasyonun, elektronik y?k ve mutasyona u?ram?? aminoasitlerin aktif enzim durumuna uzakl??? gibi baz? parametreleri de?i?tirdi?i saptand?. Bu durumlarda, pirazinamidaz?n yap? ve fonksiyonu de?i?ti ve antibiyotik etkisizdi ve sonu?ta M. tuberculosis'de pirazinamide dirence neden oldu.
Sonu?: Sonu? olarak, bu ?al??mada pirazinamide diren?li klinik M. tuberculosis izolatlar?nda protein de?i?iklikleri tan?mland?.
Anahtar Kelimeler: Mycobacterium tuberculosis, ila? direnci, pirazinamidaz, protein yap?s?, kodon.
SUMMARY
Study of pyrazinamidase structural changes in pyrazinamide resistant and susceptible isolates of Mycobacterium tuberculosis
Azam AHMADY1, Toktam POOLAD2, Poorya RAFEE1, Mojtaba TOUSHEH3, Manijeh KAHBAZI1,
Mohammad ARJOMANDZADEGAN1
1 Tuberculosis and Pediatric Infectious Diseases Research Center, Faculty of Medicine, Arak University,
Arak, Iran,
2 Department of Microbiology, Kordestan Science and Research Branch, Islamic Azad University, Arak, Iran,
3 Department of Cellular and Molecular, Isfahan University, Isfahan, Iran.
Introduction: Pyrazinamide is one of the first line four drugs for treatment of tuberculosis. It was proved that mutations in two nucleotides of 359 and 374 pnc genes are highly associated with resistance to pyrazinamide.
Matedials and Methods:In this study, mutations in these two codones in 30 clinical isolates of Mycobacterium tuberculosis were detected by means of sequencing. Protein structures encoded by this gene with and without mutation were investigated in resistant and susceptible isolates to pyrazinamide, respectively.
Results: Mutation in the positions 359 and 374 altered some parameters like change in electronic charge, distance change of mutated amino acids to situation of active enzyme and metal connection situation. In these conditions, structure and function of pyrozinamidase enzyme were changed and antibiotic was ineffective and consequently caused resistance to pyrazinamide? in M. tuberculosis.
Conclusion: This work was revealed protein changes in resistance to pyrazinamide in clinical isolates of M. tuberculosis.
Key Words: Mycobacterium tuberculosis, drug resistance, pyrazinamidase, protein structure, codon.
Tuberk Toraks 2013; 61(2): 110-114 • doi: 10.5578/tt.3888
Geli? Tarihi/Received: 22.06.2012 - Kabul Edili? Tarihi/Accepted: 03.04.2013
INTRODUCTION
Pyrazinamide (PZA) antibiotic is from the nicotinamide analogues which despite is a strong anti bacterial especially in the fairly acidic environment of body microphages and also in zones with acute inflammation. PZA is an analog of nicotinamide and a prodrug that inhibits the growth of Mycobacterium tuberculosis. PZA diffuses into M. tuberculosis cell, where the enzyme pyrazinamidase (PZase) converts PZA to the active form pyrazinoic acid (1). Pyrazinoic acid was thought to inhibit the enzyme fatty acid synthase (FAS) I, which is disrupts membrane potential and interferes with energy production, necessary for survival of M. tuberculosis it is weakly bactericidally effective on the M. tuberculosis yet, it is an efficient drug in the first two-month treatment of tuberculosis, when there are acute inflammation differences in body, and its usage has decreased the treatment duration and the possibility of relapse (3,4,5).
PZA requires conversion to the bactericidal compound pyrazinoic acid by the bacterial PZase activity of nicotinamidase to show activity against M. tuberculosis. Mutations leading to a loss of PZase activity cause PZA resistance in M. tuberculosis. Reduction in PZase activity or deletion because of mutation in pnc gene causes resistance to PZA resistance (5,6).
Accumulation of pyrazinoic acid disrupts membrane potential, necessary for survival of M. tuberculosis at an acidic site of infection. Pyrazinoic acid binds to the ribosomal protein S1 (RpsA) and inhibits trans-translation. This may explain the ability of PZA to kill dormant mycobacterium.
Although mutations in any point of pncA gene causes resistance but frequency of mutations in nucleotides of 359 and 374 has been emphasized in literature (7).
The aim of this work was comparison study of protein structure of PZase enzyme in normal and mutated forms.
Materials and Methods
Bacterial Isolates
In this study, 30 clinical isolates of M. tuberculosis sensitive and resistant to PZA, were selected by proportion method.
PCR and Sequencing
Polymerase chain reaction (PCR) by specific primers of pnc-8 (5'-GGTTGGGTGGCCGCCGGTCAG-3') and pnc-11 (5'-GCTTTGCGGCGAGCGCTCCA-3') have been accomplished by a thermocycler (Eppendorf 5332) in anealing temperature of 64?C (5).
PCR products has been sequenced by an ABI system for detection of any mutations in two nucleotides of 359 and 374.
Bioinformatics Analysis
The three dimensional (3-D) structure of the protein coded by pnc-A gene has been evaluated by an informatics investigation using Molegro Virtual Docker (MVD) software.
Results and Discussion
Molecular Study
PCR amplification on purified DNA of 30 isolates revealed 774 bp (base pair) including the entire ORF of pncA gene (400 bp) plus a part of downstream and upstream of this gene. This confirmed correct conditions and performance of PCR reaction by pnc-8 and pnc-11 primers for amplification (Figure 1).
Results from sequencing of 744-bp fragment in all isolates revealed that susceptible isolates to PZA have any mutations in nucleotides of 359 and 374. Some of resistant isolates to PZA without any mutations in these nucleotides had mutations in other nucleotides.
Protein Study
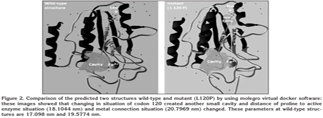
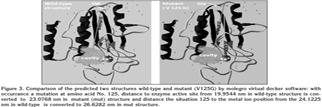
Comparing wild-type and minimized mutant structures with two codons of 120 and 125, which have respectively been created by changing T to C in 359 nucleotide and T to G in 374 nucleotide, confirms the differences in two predicted structures (Figure 2,3,4).
Mutated protein structures of changed amino acids show the importance of mutation in studied nucleotide. In nucleotide 359, after T changing to C, the Hydrophobic amino acid No. 120 (Leu) changes to amino acid with special side chain that named proline, and in nucleotide 374, after T changing to G, amino acids No. 125 [Val] changes to Gly.
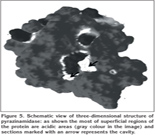
Software analysis by Molegro virtual docker showed that 3-D structure of pyrazinamidase enzyme (PZAase) consists of four alpha helix and four B-sheets (Figure 5). The special arrangement of these motifs creates a hole in the molecule that, along with Fe2+ ion, is necessary for the activity of the enzyme. This ion plays a direct, catalytic role in hydrolyze of the medicine. The important amino acid residues in performance of PZAase include residues in the enzyme active situation (D8, A134, and C138) and amino acid residues in the metal connection situation (MCS) (D49, H51, H58, and H71) (1,8). Mutation in these amino acids and other amino acids present near these situations has effect on the physicochemical activity of this enzyme. So far, in different studies, after mutation, different parameters have been attended like change in electronic charge and volume of mutated amino acids, distance change of mutated amino acids compared to situation of active enzyme and metal connection situation, and direction of side-chain of mutated amino acid (8,9).
In this paper, situation of codon 120 of three amino acids was found to be at the end of one of the helixes, and after the mutation in this protein, another small hole is created. This last one cause a distance change of proline compared to active enzyme situation and metal connection situation and direction of side-chain of mutated amino acid. But in situation of codon 125, direction of side-chain of valine changes compared to wild-type, after this amino acid changes to glycine. With relation between structure and performance of biological macromolecules, this change causes decrease or elimination of enzyme activity, and it can consequently cause resistance to pyrazinomidase (1,10). For example distance between wild type amino acid at 120 (leucine) to active site (cavity) from 17.098 nm changed to 18.1044 nm in mutant amino acid in this situation and so on.
Conclusion
In this study, the structures of wild and mutated pyrazinamidases from clinical isolates M. tuberculosis were analyzed and single amino acid replacement was studied. Mutations in nucleotides 359 and/or 374 from clinical isolates sensitive and resistant to pyrazinamidase were proved by sequencing. Bioinformatics analysis revealed that in the mutant protein structure of the pyrazinamidase enzyme was changed some parameters such as electrical charge of the mutated amino acid, the volume of the mutated amino acid, the distance of the mutated amino acid to the active site, the distance of the mutated amino acid to the metal-coordination site, and the orientation of the side-chain of the mutated amino acid. In situation of codon 125 or 120, direction of side-chain of mutated amino acid and distances to active site changes compared to wild-type. As for relation between structure and performance of enzyme, these changes causes decrease or elimination of enzyme activity, and it can consequently cause resistance to pyrazinamidase. Therefore the antibiotic will be ineffective on the bacterium.
CONFLICT of INTEREST
None declared.
REFERENCES
- Sun Z, Zhng Y. Reduced pyrazinamidase activity and the natural resistance of Mycobacterium kansasii to the antituberculosis drug pyrazinamide. Antimicrob Agents Chemother 1999; 43: 537-42.
- Boshoff? HI, Mizrahi V, Barry CE. Effects of pyrazinamide on fatty acid synthesis by whole mycobacterial cells and purified fatty acid synthase I. J Bacteriol 2002; 184: 2167-72.
- Ngo SC,? Zimhony O, Chung WJ, Sayahi H, Jacobs WR, Welchpi JT. Inhibition of isolated Mycobacterium tuberculosis fatty acid synthase I by pyrazinamide analogs. Antimicrob Agents Chemother 2007; 1: 2430-5.
- Zimhony O, Cox JS, Welch JT, Vilcheze C, Jacobs WR. Pyrazinamide inhibits the eukaryotic-like fatty acid synthetase I (FASI) of Mycobacterium tuberculosis. Nature Medicine 2000; 6: 1043-7.
- Perdiga JO, Macedo R, Malaquias A, Ferreira A, Brum L, Portugal I. Genetic analysis of extensively drug-resistant Mycobacterium tuberculosis strains in Lisbon, Portugal. Antimicrob Chemother 2010; 65: 224-7.
- Zhang H, Bi LJ, Li CY, Sun ZJ, Deng JY, Zhang ZE. Mutation found in the pncA gene of Mycobacterium tuberculosis in clinical pyrozinamide-resistant isolates from a local region of China. J Int Med Res 2009; 37: 1430-5.
- Mphahlele M, Syre H, Valvatne H, Stavrum R, Mannsaker T, Muthivhi T, et al. Pyrazinamide resistance among South African multidrug-resistant Mycobacterium tuberculosis isolates. J Clin Microbiol 2008; 46: 3459-64.
- Quiliano M, Gutierrez AH, Gilman RH, Lopez C, Evangelista W, Sotelo J, et al. Structure-activity relationship in mutated pyrazinamidases from Mycobacterium tuberculosis. Bioinformation 2011; 6: 335-9.
- Clatchy JK, Tsang AY, Cernich MS. Use of pyrazinamidase activity on Mycobacterium tuberculosis as a rapid method for determination of pyrazinamide susceptibility. Antimicrob Agents Chemother 1981; 20: 556-7.
- Zhang Y, Mitchison D. The curious characteristics of pyrazinamide.? Int J Tuberc Lung Dis 2003; 7: 6-21.
Yaz??ma Adresi (Address for Correspondence):
Dr. Mohammad ARJOMANDZADEGAN,
Tuberculosis and Pediatric Infectious
Diseases Research Center,
Arak University of Medical Sciences,
ARAK - IRAN
e-mail: mmatinam81@yahoo.com